Getting Started¶
Overview¶
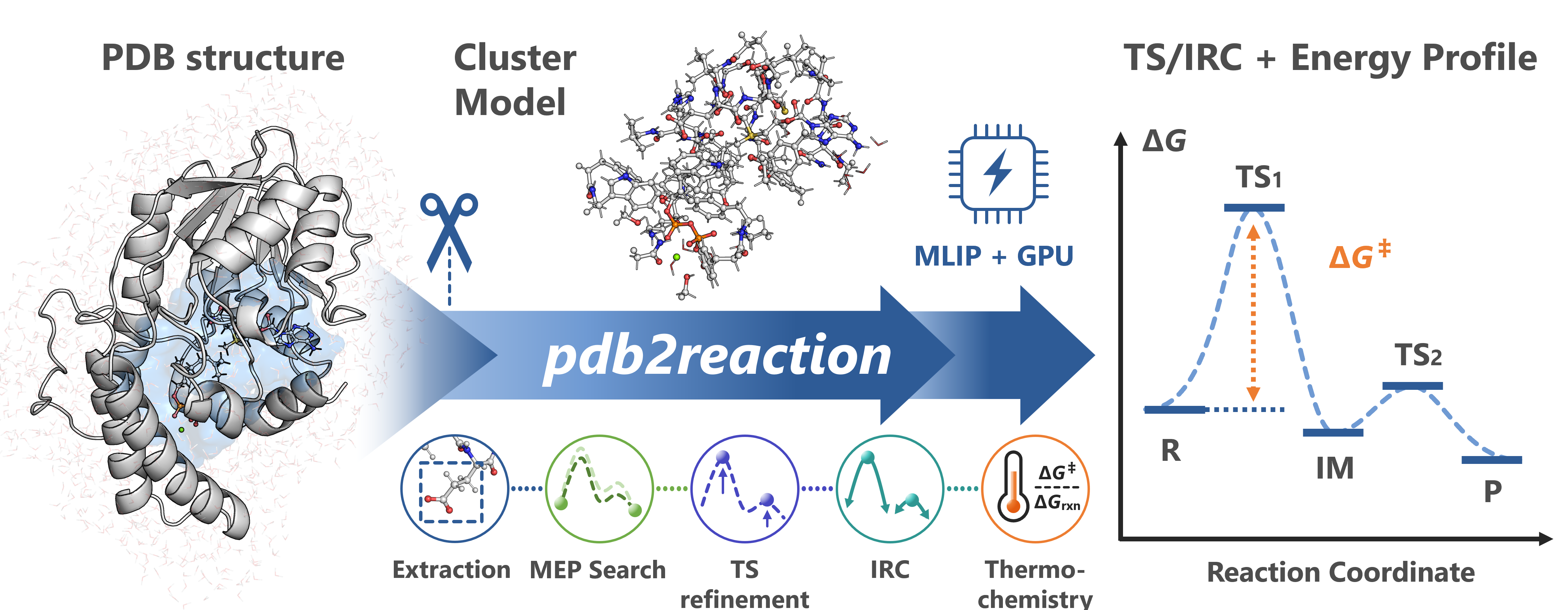
pdb2reaction is a Python CLI toolkit for modeling enzymatic reaction pathways from PDB structures using machine-learning interatomic potentials (MLIPs).
In many workflows, a single command like the one below is enough to generate a useful initial reaction path:
pdb2reaction -i 1.R.pdb 3.P.pdb -c 'SAM,GPP,MG' -l 'SAM:1,GPP:-3'
You can also run Minimum Energy Path (MEP) search → Transition State (TS) optimization → Intrinsic Reaction Coordinate (IRC) → thermochemistry → single-point DFT in a single run by adding --tsopt --thermo --dft:
pdb2reaction -i 1.R.pdb 3.P.pdb -c 'SAM,GPP,MG' -l 'SAM:1,GPP:-3' --tsopt --thermo --dft
Working examples: The
examples/directory contains completeallworkflow scripts for GPP C6-methyltransferase BezA (Tsutsumi et al., Angew. Chem. Int. Ed. 2022, 61, e202111217), covering both multi-structure MEP and scan-based pipelines.
Given (i) two or more full protein–ligand PDB files (R → … → P), or (ii) one PDB with --scan-lists/-s, or (iii) one TS candidate with --tsopt, pdb2reaction automatically:
extracts an active site model (binding pocket) around user-defined substrates to build a cluster model,
explores minimum-energy paths (MEPs) with path optimization methods such as the Growing String Method (GSM) and Direct Max Flux (DMF),
optionally optimizes transition states, runs vibrational analysis, IRC calculations, and single-point DFT calculations.
Calculations use machine-learning interatomic potentials (MLIPs). The default backend is Meta’s UMA, but ORB, MACE, and AIMNet2 are also supported via -b/--backend. Typical use cases include:
Trial-and-error exploration of reaction mechanisms at a scale where DFT-level verification would be too slow
Generating initial geometries (reactant/TS/product cluster models) for subsequent quantum-chemistry refinement
High-throughput screening of reaction pathways across substrate variants or enzyme mutants
The CLI generates multi-step enzymatic reaction mechanisms with minimal manual setup. The same workflow also works for small-molecule systems. When you skip active site model extraction (omit --center/-c and --ligand-charge/-l), you can also use .xyz or .gjf inputs.
On HPC clusters or multi-GPU workstations, pdb2reaction can scale to large cluster models (and optionally full protein–ligand complexes) by parallelizing UMA inference across nodes. Set workers and workers_per_node to enable multi-worker inference; see MLIP Calculator for configuration details. Alternative backends (ORB, MACE, AIMNet2) can be selected with -b/--backend.
Pipeline overview¶
The all subcommand runs the following stages automatically:
PDB (R, P)
|
v
[extract] Active site model extraction (cluster model)
|
v
[scan] (optional, --scan-lists/-s) Staged distance-restraint scans
|
v
[path-search] MEP search (recursive path-search, default; --refine-path False switches to path-opt)
|
v
[tsopt] TS optimization (RS-I-RFO; Dimer as alternative)
|
v
[irc] Intrinsic Reaction Coordinate
|
v
[freq] Vibrational analysis + thermochemistry (R, TS, P)
|
v
[dft] Single-point DFT energy (optional, --dft)
Each stage can also be run as a standalone subcommand. The all command orchestrates them and produces a unified summary.json and summary.log.
Key output files¶
File |
Description |
|---|---|
|
Machine-readable results (barriers, energies, bond changes, environment) |
|
Human-readable text summary with directory tree |
|
IRC-optimized R/TS/P structures per reaction step |
|
Merged MEP trajectory viewable in PyMOL/VMD |
|
Energy profile plots (electronic / Gibbs-corrected) |
Important
Input PDB files must already contain hydrogen atoms.
When you provide multiple PDBs, they must contain the same atoms in the same order (only coordinates may differ); otherwise an error is raised.
Tip
For symptom-first diagnosis, start with Common Error Recipes. If you encounter an error during setup or runtime, refer to Troubleshooting.
CLI conventions¶
Convention |
Example |
Notes |
|---|---|---|
Residue selectors |
|
Quote multi-value strings to prevent shell expansion |
Charge mapping |
|
Colon separates name and charge; comma separates entries |
Atom selectors |
|
Delimiters: space, comma, slash, backtick, backslash |
For full details, see CLI Conventions.
Recommended tools for hydrogen addition¶
If your PDB lacks hydrogen atoms, use one of the following tools before running pdb2reaction:
Tool |
Example Command |
Notes |
|---|---|---|
reduce (Richardson Lab) |
|
Fast, widely used for crystallographic structures |
pdb2pqr |
|
Adds hydrogens and assigns partial charges |
Open Babel |
|
General-purpose cheminformatics toolkit |
PyMOL |
|
Molecular visualization tool with hydrogen addition |
tleap (AmberTools) |
|
Amber force-field preparation tool |
To ensure identical atom ordering across multiple PDB inputs, apply the same hydrogen-addition tool with consistent settings to all structures.
Quickstart routes (recommended)¶
For setup and dependency installation, see Installation.
Command line basics¶
The main entry point is the pdb2reaction command, installed via pip. A shorthand alias p2r is also registered by the pdb2reaction package (same setuptools entry point; you get both after pip install pdb2reaction) — all commands can be run with either name. Internally it uses the Click library, and the default subcommand is all.
That means:
pdb2reaction [OPTIONS]...
# is equivalent to
pdb2reaction all [OPTIONS]...
The all command runs the full pipeline—cluster extraction, MEP search, TS optimization, vibrational analysis, and optional DFT—in a single invocation.
All high-level workflows share two important options when you use cluster extraction:
-i/--input: one or more full structures (reactant, intermediate(s), product).-c/--center: how to define the substrate / extraction center (e.g., residue names or residue IDs).
If you omit --center/-c, cluster extraction is skipped and the full input structure is used directly.
Main workflow modes¶
Mode |
Description |
Quickstart |
|---|---|---|
Multi-structure MEP (≥ 2 PDBs) |
Take full PDBs along a putative reaction coordinate (R → … → P), extract cluster models, run recursive MEP search, and optionally refine with TS / IRC / freq / DFT per segment. |
|
Single-structure + staged scan (1 PDB + |
Drive one PDB through staged distance-restraint scans on the cluster model; each stage seeds the recursive |
|
Single-structure TSOPT-only (1 PDB + |
Skip MEP entirely; optimize a TS candidate, run IRC in both directions, optimize endpoints, and optionally run freq / DFT on R/TS/P. |
Important
Single-input runs require either --scan-lists/-s (staged scan → GSM) or --tsopt (TSOPT-only). Supplying only a single -i without one of these will not trigger a full workflow.
Important CLI options and behaviors¶
Below are the most commonly used options across workflows.
Option |
Description |
|---|---|
|
Input structures. ≥ 2 PDBs → MEP search; 1 PDB + |
|
Defines the substrate / extraction center. Supports residue names ( |
|
Charge info: mapping ( |
|
Hard override of net system charge. |
|
Spin multiplicity (e.g., |
|
Enable TS optimization and IRC. |
|
Select MLIP backend ( |
For the complete option matrix, see CLI Conventions and the generated CLI reference.
Run summaries¶
Every pdb2reaction all run writes:
summary.log– text summary, andsummary.json– JSON summary.
They typically contain:
the exact CLI command invoked,
global MEP statistics (e.g. maximum barrier, path length),
per-segment barrier heights and key bond changes,
energies from the MLIP backend, thermochemistry, and DFT post-processing (where enabled).
Each segment directory under path_search/ (or path_opt/ when --refine-path False is used) also gets its own summary.log and summary.json, so you can inspect local refinements independently.
CLI commands¶
Most users will primarily call pdb2reaction all. The CLI also exposes individual subcommands; each supports -h/--help and (for the calculation/scan/extract/utility commands) --help-advanced for the full list. For the subcommand index with per-command documentation links, see the documentation home.
Agent Skills¶
pdb2reaction ships AI-agent instructions under .claude/skills/
covering the CLI subcommands, structure I/O (PDB / XYZ / GJF), backend
installation (UMA / Orb / MACE / AIMNet2 / DFT / xtb), canonical
workflows, output parsing, and HPC operation. Copy .claude/skills/
into your project repository or ~/.claude/skills/ for agent platforms
like Claude Code or Cursor.
Getting help¶
For any subcommand:
pdb2reaction <subcommand> --help
pdb2reaction <subcommand> --help-advanced
pdb2reaction all --help-advanced
For detailed MLIP backend options, see MLIP Calculator.
If you encounter any issues, please open an Issue on the GitHub repository.