pdb2reaction Documentation¶
Version: v0.3.7
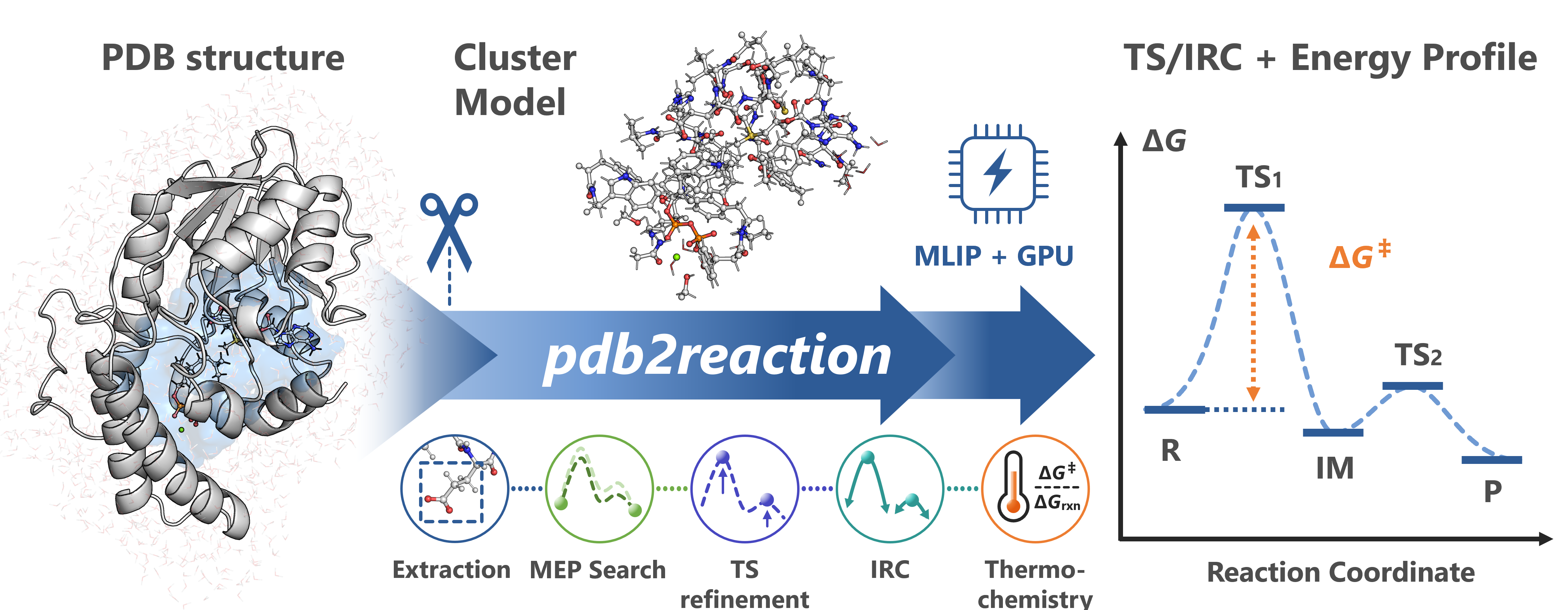
pdb2reaction is a Python CLI toolkit for automated enzymatic reaction-path modeling directly from PDB structures using machine-learning interatomic potentials (MLIPs).
Start here¶
Goal |
Workflow |
|---|---|
First end-to-end run |
|
Reactant only |
|
TS candidate available |
|
Run failure / error |
Refer to Installation for prerequisites.
Subcommands¶
Subcommand |
Description |
|---|---|
End-to-end workflow: extraction → scan → MEP → TS optimization → IRC → thermochemistry → DFT |
|
Extract active site model (binding pocket) from protein–ligand complex |
|
Resolve PDB alternate locations |
|
Repair PDB element columns (77–78) |
|
Single-structure geometry optimization (L-BFGS or RFO. [+ optional flatten]) |
|
Transition state optimization (Dimer or RS-I-RFO. [+ optional flatten]) |
|
Single-step MEP optimization via GSM or DMF (from 2 structures) |
|
Recursive multi-step MEP search with automatic refinement (2+ structures) |
|
1D bond-length driven scan with restraints |
|
2D distance grid scan |
|
3D distance grid scan |
|
Vibrational frequency analysis & thermochemistry |
|
Intrinsic Reaction Coordinate calculation |
|
Single-point DFT calculations (GPU4PySCF / PySCF) |
|
Plot energy profiles from XYZ trajectories |
|
Draw an energy diagram from numeric values |
|
Detect and report covalent bond changes between consecutive structures |
Configuration & Reference¶
Topic |
Page |
|---|---|
Getting Started |
|
CLI conventions & input requirements |
|
Cluster boundary atoms (link hydrogens, |
|
Common errors & fixes |
|
CLI command reference |
|
YAML configuration options |
|
MLIP backend settings |
|
Terminology |
System Requirements¶
Hardware¶
OS: Linux
GPU: CUDA 12.x compatible
VRAM: 8 GB+ recommended
RAM: 16 GB+ recommended
Software¶
Python >= 3.11
PyTorch with CUDA support
CUDA 12.x toolkit
For setup, see Installation.
Agent Skills¶
pdb2reaction ships AI-agent instructions under .claude/skills/ covering CLI subcommands, structure I/O, backend installation, workflows, output parsing, and HPC operation. See .claude/skills/README.md for the full skill index and installation instructions.
Citation¶
A preprint describing pdb2reaction is in preparation. Currently, if you find this work helpful for your research, please cite the software itself:
@software{ohmura2026pdb2reaction,
author = {Ohmura, Takuto},
title = {pdb2reaction},
year = {2026},
month = {4},
version = {0.3.6},
url = {https://github.com/t-0hmura/pdb2reaction},
license = {GPL-3.0},
doi = {10.5281/zenodo.19197865}
}
License¶
pdb2reaction is distributed under the GNU General Public License version 3 (GPL-3.0).
Getting Help¶
# General help
pdb2reaction --help
# Command help
pdb2reaction <subcommand> --help
# Advanced options (dry-run, internal tuning, etc.)
pdb2reaction <subcommand> --help-advanced
For issues and feature requests, visit the GitHub repository.